Gene expression is not controlled by a single switch. Instead, it is regulated through complex systems known as regulatory networks. These networks coordinate how genes are activated, suppressed, or modified so that cells can function properly.
Regulatory networks are essential for processes such as development, adaptation to environmental changes, immune responses, and cellular specialization. By controlling when and how genes are expressed, cells maintain balance and respond to internal and external signals.
Understanding gene regulatory networks helps scientists explain how biological systems maintain stability while remaining flexible enough to adapt.
What Is Gene Expression?
Gene expression is the process by which the information stored in DNA is used to produce functional products such as proteins or RNA molecules.
This process occurs in two primary stages:
- Transcription – DNA is copied into messenger RNA (mRNA).
- Translation – mRNA is used as a template to build proteins.
However, cells rarely express genes randomly. Instead, regulatory systems determine:
- When a gene should be activated
- How much protein should be produced
- In which cell types a gene should function
- How long gene activity should continue
These decisions are governed by regulatory networks.
What Are Gene Regulatory Networks?
Gene regulatory networks (GRNs) are interconnected systems of genes, proteins, and regulatory elements that control gene expression.
In these networks:
- Some genes produce proteins that regulate other genes.
- Regulatory proteins activate or repress transcription.
- Feedback loops maintain stability in gene activity.
A regulatory network can involve hundreds or thousands of genes interacting with one another.
These networks allow cells to coordinate complex behaviors rather than relying on isolated genetic signals.
Components of Gene Regulatory Networks
Several key biological components form the structure of gene regulatory networks.
Regulatory Genes
Regulatory genes produce proteins that control the activity of other genes.
These proteins often function as transcription factors that bind to DNA and influence whether transcription begins.
Transcription Factors
Transcription factors are specialized proteins that recognize specific DNA sequences near genes.
Their functions include:
- Activating gene transcription
- Repressing gene transcription
- Coordinating multiple genes simultaneously
A single transcription factor may regulate dozens or even hundreds of target genes.
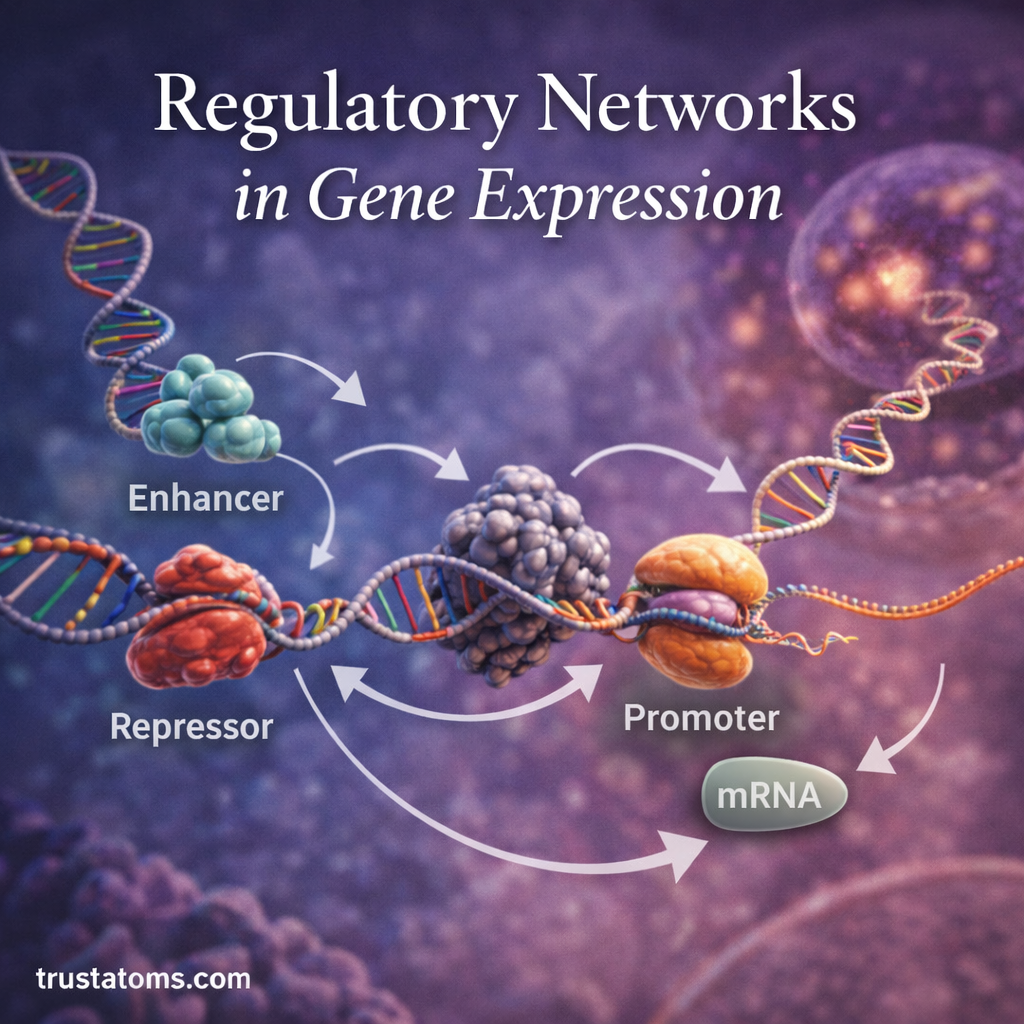
Promoters
Promoters are DNA regions located near the start of genes.
They act as binding sites for transcription machinery and transcription factors.
Promoters determine whether a gene can begin transcription.
Enhancers and Silencers
Enhancers and silencers are regulatory DNA sequences that influence gene expression from a distance.
Enhancers increase transcription activity, while silencers decrease or suppress gene expression.
These regulatory elements help fine-tune gene activity.
Types of Regulatory Interactions
Gene regulatory networks operate through different types of interactions.
Activation
Activation occurs when a regulatory protein increases the likelihood that transcription will begin.
This often happens when a transcription factor binds to enhancer regions.
Repression
Repression happens when regulatory proteins block transcription.
Repressors may bind to DNA sequences that prevent RNA polymerase from initiating transcription.
Cooperative Regulation
In many cases, multiple transcription factors work together.
Cooperative regulation allows cells to integrate multiple signals before activating a gene.
For example, a gene may only be expressed when two different transcription factors bind simultaneously.
Feedback Loops in Gene Regulatory Networks
Feedback loops are critical features of regulatory networks. They help maintain stability and prevent uncontrolled gene expression.
Two major types of feedback loops exist.
Positive Feedback
Positive feedback amplifies gene activity.
When a gene activates its own expression indirectly, the result can be sustained gene activity.
Positive feedback is often involved in:
- Cell differentiation
- Developmental pathways
- Signal amplification
Negative Feedback
Negative feedback reduces gene activity after a certain threshold is reached.
This type of regulation prevents excessive protein production and maintains cellular balance.
Negative feedback is common in:
- Hormone regulation
- Metabolic pathways
- Homeostasis systems
Regulatory Networks in Cell Differentiation
One of the most important roles of gene regulatory networks is controlling cell differentiation.
All cells in a multicellular organism contain the same DNA, but regulatory networks determine which genes are active in each cell type.
Examples include:
- Neurons activating genes for neurotransmitter signaling
- Muscle cells activating genes for contraction proteins
- Immune cells activating genes related to pathogen defense
Regulatory networks establish stable patterns of gene expression that define cell identity.
Environmental Influence on Gene Regulation
Gene regulatory networks are highly responsive to environmental signals.
Cells constantly monitor their surroundings and adjust gene expression accordingly.
Environmental signals that influence gene regulation include:
- Nutrient availability
- Temperature changes
- Chemical signals
- Stress conditions
- Hormones and signaling molecules
Signal transduction pathways transmit these signals to the nucleus, where transcription factors alter gene activity.
This allows organisms to adapt quickly to changing conditions.
Regulatory Networks in Development
During development, gene regulatory networks guide the formation of tissues and organs.
Specific networks activate groups of genes in precise sequences.
These developmental regulatory systems control:
- Embryonic pattern formation
- Tissue specialization
- Organ development
- Body axis formation
Small changes in developmental regulatory networks can produce significant changes in organism structure, which plays a role in evolution.
Gene Regulatory Networks and Disease
Disruptions in gene regulatory networks can lead to disease.
When regulatory pathways malfunction, genes may become overactive or silenced when they should not be.
Examples include:
Cancer
Many cancers arise from mutations in regulatory genes or transcription factors.
These mutations can cause uncontrolled cell division by activating growth genes or disabling tumor suppressor genes.
Genetic Disorders
Some inherited disorders occur when regulatory elements controlling gene expression are damaged.
Even if the gene itself is normal, faulty regulation can disrupt normal biological processes.
Neurological Conditions
Certain neurological disorders are linked to altered gene regulation during brain development.
Improper regulatory signaling can affect neuron growth and connectivity.
How Scientists Study Regulatory Networks
Researchers use several technologies to analyze gene regulatory networks and understand how genes interact.
Common methods include:
- RNA sequencing (RNA-seq) to measure gene expression levels
- ChIP-seq to identify transcription factor binding sites
- CRISPR gene editing to test regulatory gene functions
- Single-cell sequencing to analyze gene regulation in individual cells
These tools allow scientists to map entire regulatory networks across genomes.
Final Thoughts
Regulatory networks in gene expression form the foundation of cellular control systems. Instead of acting independently, genes interact through complex networks of regulatory proteins and DNA elements that coordinate biological activity.
These networks allow cells to adapt, differentiate, and maintain internal balance. From embryonic development to immune responses, gene regulatory systems guide many of the processes that sustain life.
As research advances, scientists continue uncovering how regulatory networks shape biological complexity and influence health and disease.