Chromatin remodeling is one of the most important processes controlling how genes are turned on or off inside living cells. While DNA contains the instructions for building proteins, not every gene is active at the same time. Cells rely on complex regulatory systems to control access to DNA, and chromatin remodeling is a central part of that regulation.
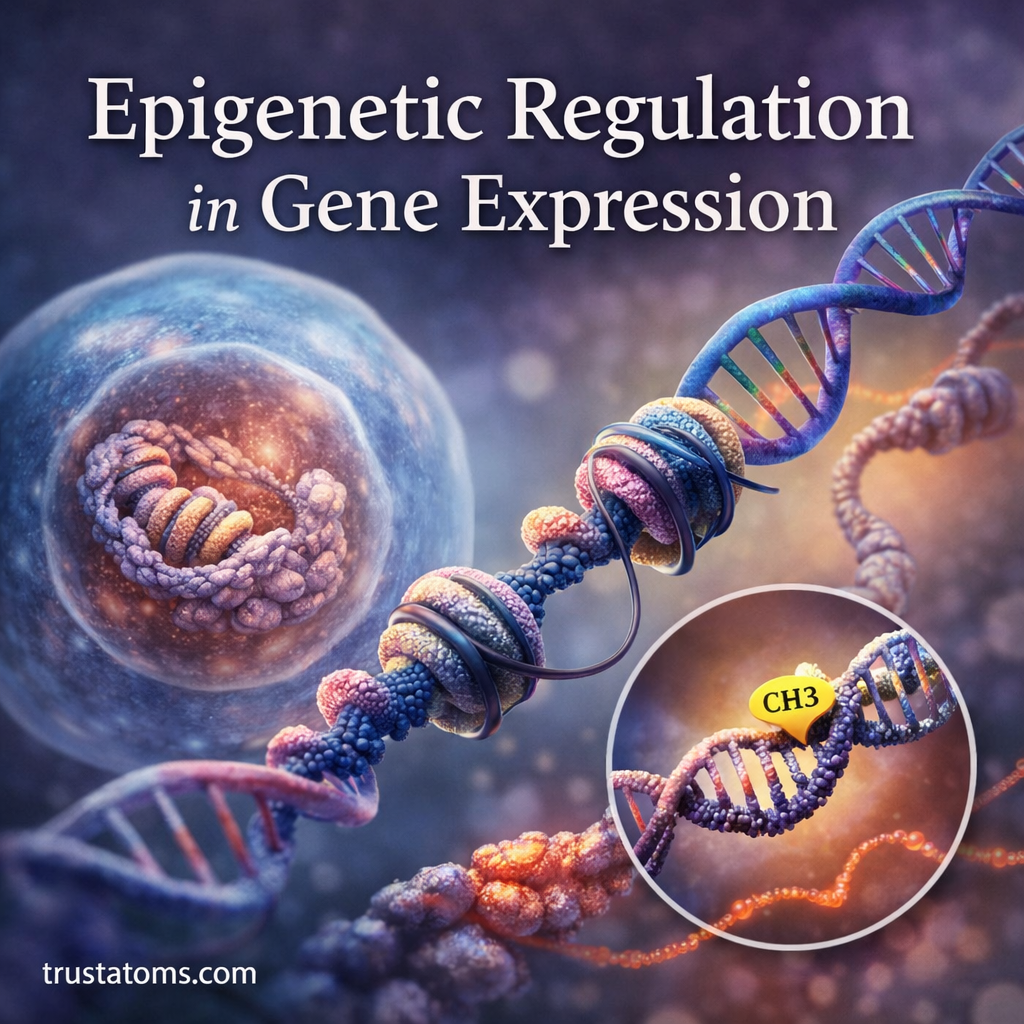
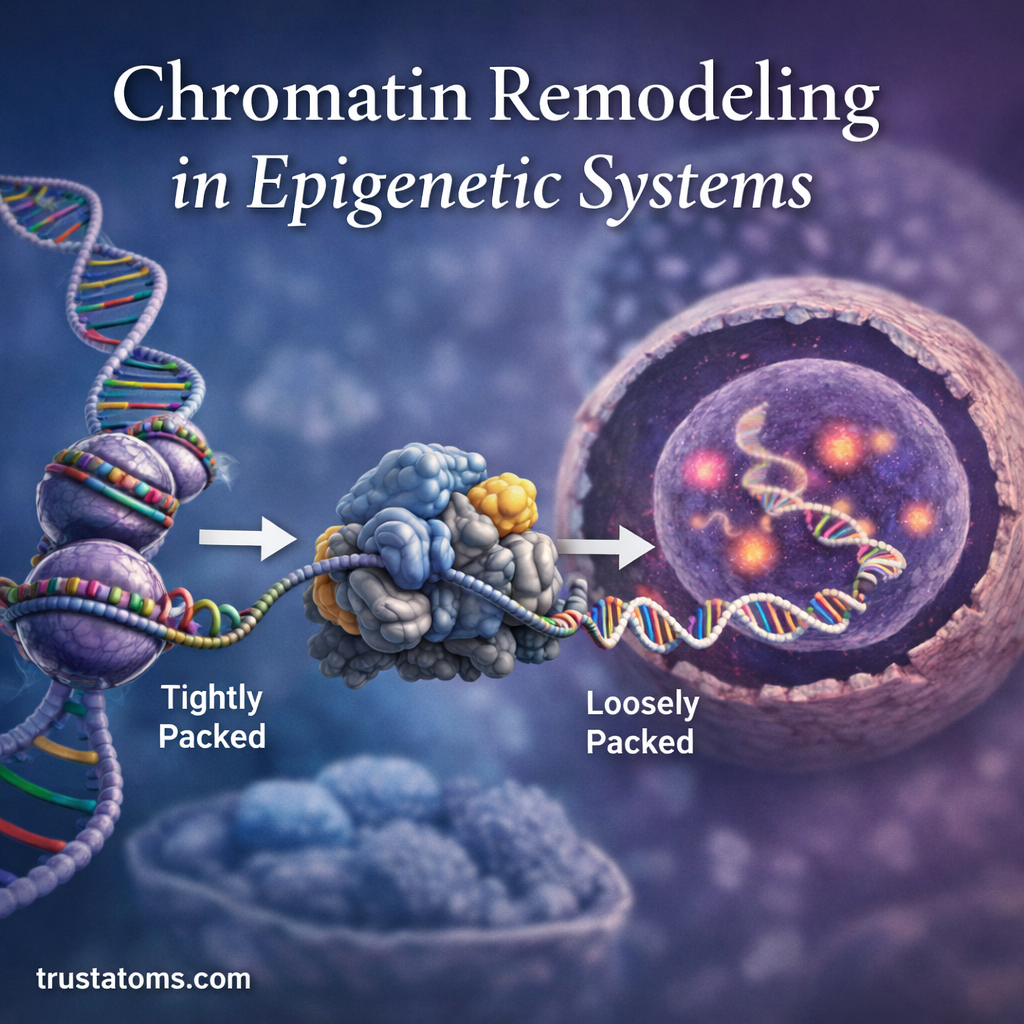
In epigenetic systems, chromatin remodeling alters the physical structure of DNA and its associated proteins without changing the DNA sequence itself. By shifting how tightly DNA is packaged, cells can determine which genes are accessible for transcription and which remain silent.
Understanding chromatin remodeling helps explain how identical DNA can produce many different cell types, how organisms respond to environmental changes, and how certain diseases develop.
What Is Chromatin?
Chromatin is the structural complex formed by DNA wrapped around proteins called histones. In eukaryotic cells, DNA is extremely long, and chromatin allows it to be efficiently packaged inside the cell nucleus.
Chromatin has two major components:
- DNA molecules containing genetic instructions
- Histone proteins that organize and compact DNA
DNA winds around groups of histone proteins to form structures known as nucleosomes. Each nucleosome consists of about 147 base pairs of DNA wrapped around a histone core.
This packaging system allows meters of DNA to fit inside a microscopic nucleus while still permitting controlled access to specific genes.
Chromatin States: Open vs Closed
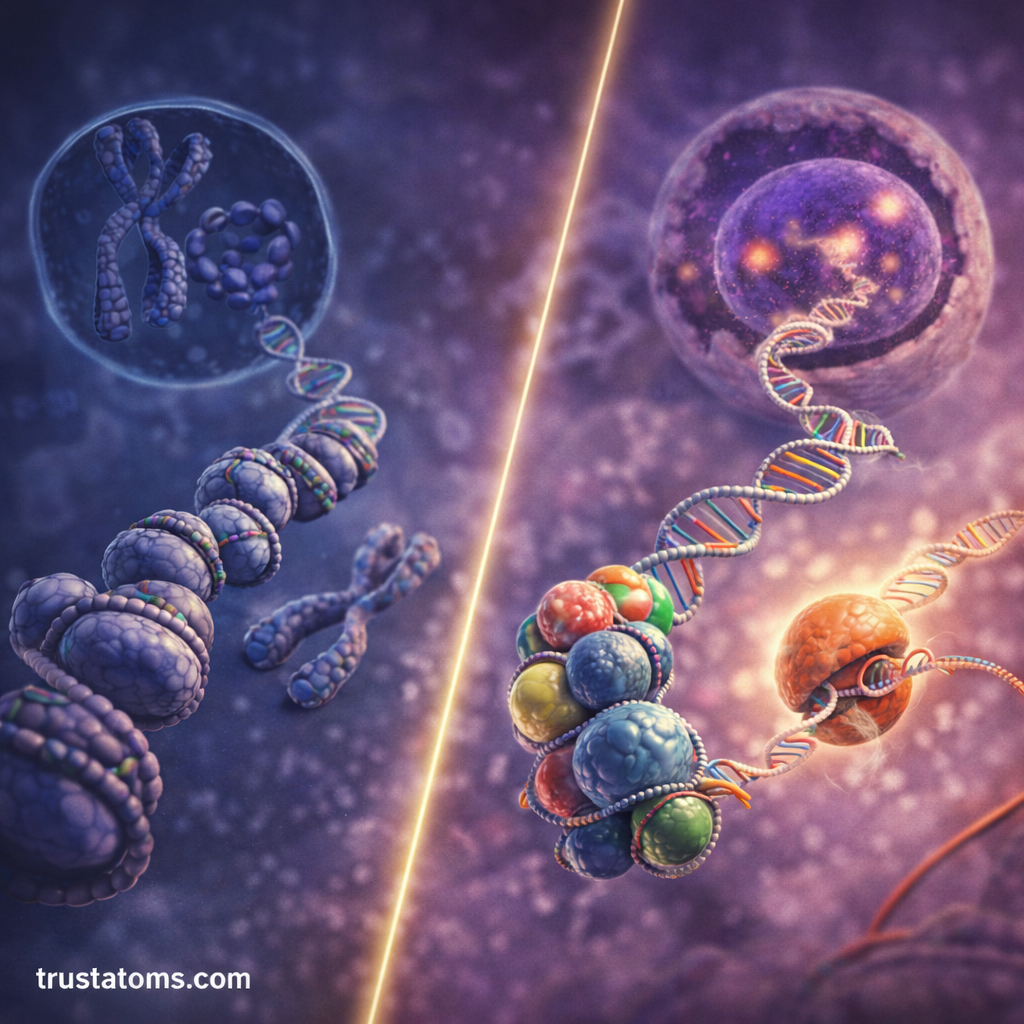
Chromatin does not remain in a single static form. Instead, it can shift between different structural states that influence gene activity.
Two major chromatin states exist:
Euchromatin
Euchromatin is loosely packed chromatin that allows genes to be easily accessed by transcription machinery.
Characteristics include:
- Active gene transcription
- Loosely wrapped DNA
- High accessibility to transcription factors and RNA polymerase
Genes located in euchromatin regions are generally active or ready to be activated.
Heterochromatin
Heterochromatin is tightly packed chromatin that prevents transcription machinery from reaching the DNA.
Key features include:
- Gene silencing
- Dense DNA packing
- Limited accessibility to regulatory proteins
Heterochromatin is commonly found in regions that must remain inactive, such as repetitive DNA sequences or structural chromosome areas.
Chromatin remodeling allows cells to switch between these two states depending on regulatory needs.
What Is Chromatin Remodeling?
Chromatin remodeling refers to the process by which cells change the structure or position of nucleosomes to control DNA accessibility.
Instead of altering the DNA code, chromatin remodeling modifies how DNA is packaged.
This process may involve:
- Sliding nucleosomes along DNA
- Removing nucleosomes
- Replacing histones with variants
- Loosening or tightening DNA wrapping
These changes determine whether specific genes can be transcribed.
Chromatin remodeling is essential for many biological processes, including cell differentiation, development, and response to environmental signals.
Chromatin Remodeling Complexes
Chromatin remodeling is carried out by specialized protein machines known as chromatin remodeling complexes.
These complexes use energy from ATP to physically reposition nucleosomes.
Major families of chromatin remodeling complexes include:
SWI/SNF Family
The SWI/SNF complexes help open chromatin structures by moving nucleosomes away from promoter regions, making DNA accessible for transcription.
They are heavily involved in gene activation and cellular differentiation.
ISWI Family
ISWI complexes regulate nucleosome spacing and help maintain organized chromatin structure across the genome.
They contribute to both gene activation and repression depending on context.
CHD Family
CHD (Chromodomain Helicase DNA-binding) complexes participate in transcription regulation and developmental gene control.
They often recognize specific histone modifications before remodeling chromatin.
INO80 Family
INO80 complexes are involved in:
- DNA repair
- Replication
- Chromosome stability
These complexes help reposition nucleosomes around damaged DNA regions.
Together, these remodeling complexes constantly adjust chromatin architecture throughout the cell’s life.
Histone Modifications and Remodeling
Chromatin remodeling works closely with chemical changes to histone proteins known as histone modifications.
These modifications act like molecular signals that influence chromatin structure.
Common histone modifications include:
- Acetylation
- Methylation
- Phosphorylation
- Ubiquitination
Each modification affects chromatin in different ways.
Histone Acetylation
Histone acetylation generally promotes gene activation.
Acetyl groups added to histone tails reduce the attraction between DNA and histones, loosening chromatin structure and allowing transcription machinery to bind DNA.
Histone Methylation
Histone methylation can either activate or repress gene expression depending on the location and number of methyl groups added.
Certain methylation patterns mark genes for activation, while others mark regions for long-term silencing.
Chromatin remodeling complexes often recognize these chemical tags and adjust nucleosome positions accordingly.
Role of Chromatin Remodeling in Epigenetics
Epigenetics refers to heritable changes in gene activity that occur without altering the DNA sequence.
Chromatin remodeling is one of the primary mechanisms that enables epigenetic regulation.
Epigenetic control systems allow cells to:
- Maintain stable patterns of gene expression
- Respond to environmental signals
- Guide developmental processes
For example, during embryonic development, chromatin remodeling helps activate genes necessary for forming different tissues.
Muscle cells, neurons, and skin cells all share the same DNA, but chromatin remodeling determines which genes each cell type expresses.
Chromatin Remodeling in Development
During development, cells transition from a pluripotent state to specialized cell types.
Chromatin remodeling helps establish these identities by activating lineage-specific genes while silencing unrelated pathways.
Examples include:
- Neural genes activated in developing neurons
- Muscle genes activated in muscle tissue
- Immune genes activated in white blood cells
Once established, many of these chromatin patterns remain stable through cell divisions, allowing tissues to maintain their specialized functions.
Chromatin Remodeling and Environmental Influence
Environmental conditions can also influence chromatin remodeling.
Factors such as:
- Nutrition
- Stress
- Toxins
- Temperature
- Physical activity
can alter epigenetic markers and chromatin accessibility.
These changes may temporarily affect gene expression or, in some cases, persist across multiple cell generations.
This connection between environment and gene regulation is an important area of epigenetic research.
Chromatin Remodeling and Disease
Disruptions in chromatin remodeling systems are linked to numerous diseases.
Because chromatin structure controls gene expression, errors in remodeling can activate harmful genes or silence protective ones.
Examples include:
Cancer
Mutations in chromatin remodeling genes are common in many cancers.
For example:
- SWI/SNF complex mutations appear in several tumor types
- Improper chromatin regulation can activate oncogenes or disable tumor suppressor genes
Neurological Disorders
Chromatin remodeling abnormalities are also associated with:
- Autism spectrum disorders
- Intellectual disabilities
- Neurodevelopmental syndromes
These conditions often arise when gene expression patterns in brain development are disrupted.
Aging
Chromatin structure gradually changes with age, altering gene regulation and cellular function.
Epigenetic drift over time may contribute to age-related diseases.
How Scientists Study Chromatin Remodeling
Modern molecular biology techniques allow researchers to study chromatin structure across the genome.
Key methods include:
- ChIP-seq (Chromatin Immunoprecipitation Sequencing) for detecting histone modifications
- ATAC-seq for measuring DNA accessibility
- MNase-seq for mapping nucleosome positions
- Hi-C sequencing for analyzing chromatin folding in three dimensions
These technologies provide insights into how chromatin architecture controls gene regulation at a genome-wide scale.
Final Thoughts
Chromatin remodeling is a central mechanism in epigenetic systems that determines how DNA instructions are interpreted inside cells. By shifting the structure and accessibility of chromatin, cells can precisely regulate gene expression without altering the genetic code itself.
This process plays a critical role in development, cellular identity, environmental response, and disease prevention. As research advances, scientists continue uncovering how chromatin remodeling shapes biological complexity and influences health throughout life.
Understanding chromatin remodeling not only deepens our knowledge of genetics but also opens new possibilities for medical therapies that target epigenetic regulation.